EPI2ME Labs 22.05.01 Release
Dear Nanopore Community,
We are pleased to release a new version of the EPI2ME Labs software that introduces an updated and simplified user experience. EPI2ME Labs is now a standalone application that continues to provide a clean interface to accessing bioinformatics workflows and tutorials for the analysis of nanopore sequence data. This release also brings our workflow capabilities to users working from Windows computers.
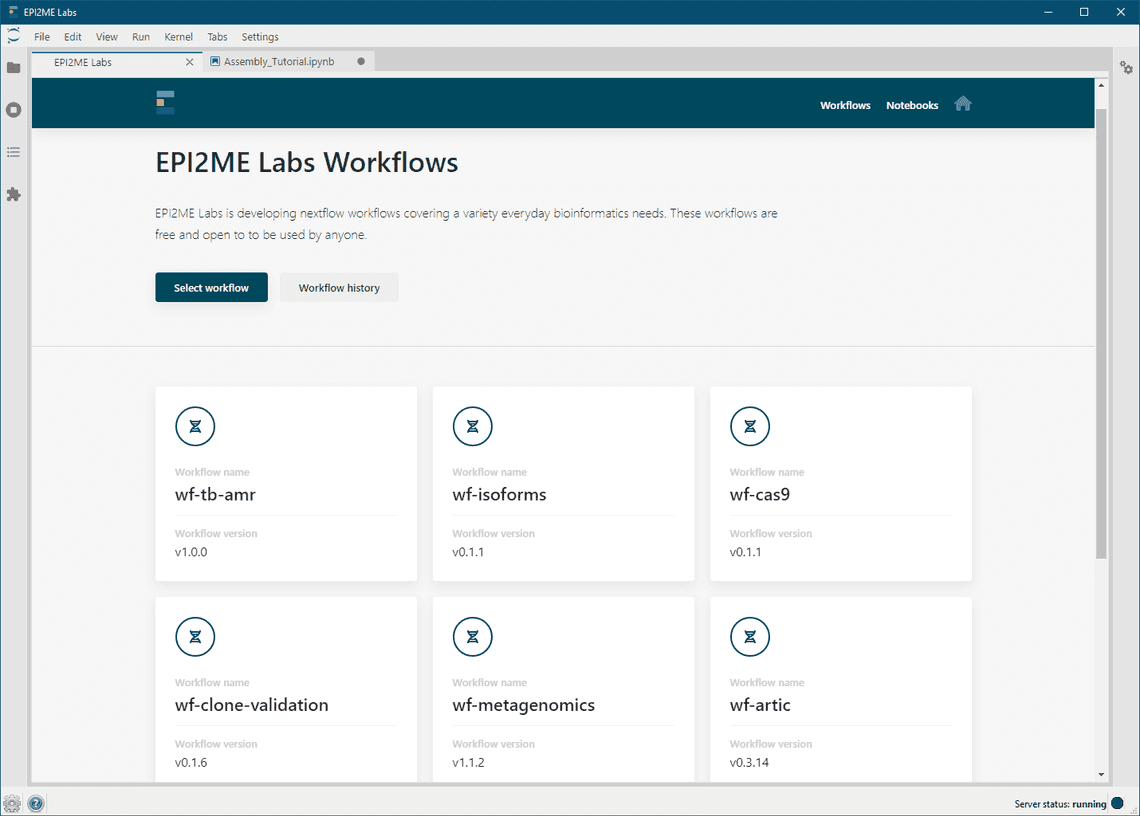
There are a number of key changes within the updated EPI2ME Labs software that are summarised below:
The EPI2ME Labs Launcher software has been deprecated - the new EPI2ME Labs software automatically orchestrates the download, start and stop of the required Docker containers. The EPI2ME Labs software functions as a standalone application and now delivers the Jupyter interface itself. The EPI2ME Labs software is focusing on our collection of Nextflow based workflows. These have been improved with additional in-application documentation to better describe the workflows, their parameters, and configuration. Updates to the workflow integration make it simpler to re-run workflows that have been pre-configured with preferred parameters, databases, or reference genomes. The command-line access to the software has been maintained and for advanced users, the ability to start multiple sessions on a server remains possible. The installer software has been updated to simplify the installation for Windows users - the installer scripts now orchestrate the installation of most dependencies (WSL2, an Ubuntu VM, Nextflow and its Java dependencies). The only unmanaged software dependency is now Docker, which can be downloaded as Docker Desktop.
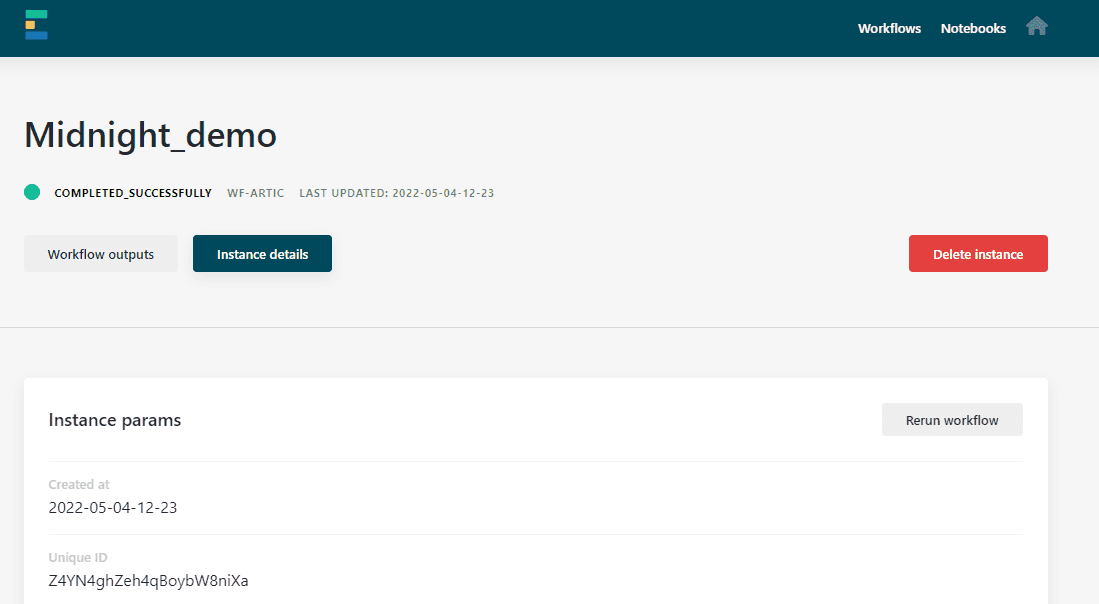
The EPI2ME Labs software is available from our Downloads page. Additional information is provided in the project’s CHANGELOG.
It is recommended that a previous installation of the EPI2ME Labs Launcher is first removed - the new EPI2ME Labs software cannot uninstall the now deprecated EPI2ME Labs Launcher software.
New workflow releases
The EPI2ME Labs release also includes updates to some of our workflows. The template for our Nextflow based workflows can be found at wf-template.
Notably wf-tb-amr has been updated to version v1.0.0. This version now uses the bcftools mpileup software for variant calling and uses a larger collection of WHO defined variants in antibiotic resistance association. Please see the changelog for additional information on the changes. wf-metagenomics has been updated to version v1.1.3. This update includes in-application documentation improvements required for functionality with the EPI2ME Labs software.
We look forwards to any feedback and would welcome recommendations for new workflows, tutorials, and features.
Related Posts
Information

