EPI2ME Labs 22.06.01 Release
Dear Nanopore Community,
We are pleased to release a new infectious diseases workflow and a collection of minor updates to our EPI2ME Labs product.
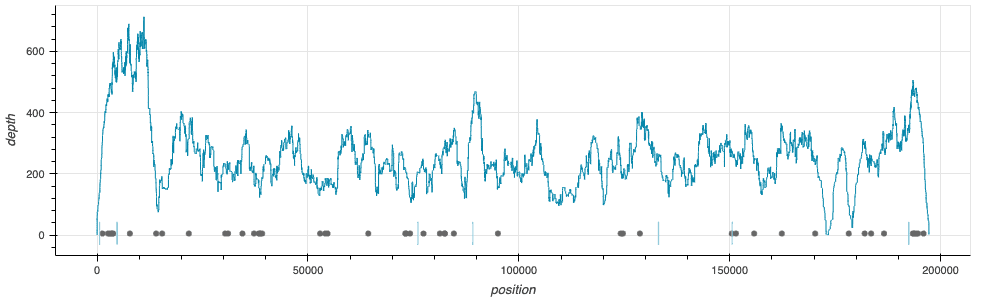
Release of a new workflow, wf-mpx, with initial version v0.0.1 for the basic analysis of sequence data prepared from samples containing monkeypox virus. The workflow is now available through the EPI2ME Labs application. An article describing the workflow, and providing some recommendations for how the mapped sequence data might best be considered, can be found in a separate Monkeypox Analysis post.
- The wf-artic software for analysis of SARS-CoV-2 sequence data has been updated to v0.3.16. This update includes an updated primer scheme for the NEB VarSkip V2b design. There is also a change that allows for the passing of additional parameters to the Pangolin software component - further details may be found in the project’s CHANGELOG.
- Several of the tutorials included in EPI2ME Labs have been deprecated. Tutorials are intended to introduce data analysis concepts but the Jupyter notebook is not an ideal vehicle for the production analysis of sequence data. For production analysis of Nanopore sequence data we recommend only our Nextflow-based workflows - these are simpler to run, have richer analysis reports and are more actively maintained and developed. A separate post explains more about the rationale behind this decision and lists the tutorials that have been deprecated.
- The EPI2ME Labs file browser has been updated and simplifies connecting your sequence data to a workflow. A collection of minor fixes have also been included in the update. These changes have been implemented in the epi2melabs-notebook v1.1.21 container.
- Our wf-isoforms workflow (v0.1.3) has been updated to use the latest version of pychopper (v2.7.0) that now supports the SQK-PCS111 kit. Please see the CHANGELOG for additional information.
We welcome any feedback and would love to hear recommendations for workflow or interface improvements.
Related Posts
Information

