Nanopore-only T2T assembly of a human genome

We are pleased to announce the release of a new addition to the Oxford Nanopore Open Data project as part of London Calling 2024: a Nanopore-only telomere-to-telomere (T2T) assembly dataset. This dataset was created using our new telomere-to-telomere (T2T) workflow, combining ultra-long reads, Pore-C and our new assembly-polishing chemistry to completely resolve haplotypes and achieve a state-of-the-art Q50 human assembly. To register your interest in the T2T workflow, please click here.
The following cell line samples were obtained from the NIGMS Human Genetic Cell Repository at the Coriell Institute for Medical Research: GM24385
This dataset contains the inputs and outputs of the T2T workflow, comprising basecalled reads from sequencing runs using four PromethION flow cells:
- Two Ultra Long (SQK-ULK114) runs, basecalled using the model
dna_r10.4.1_e8.2_400bps_sup@v5.0.0 - One Pore-C (SQK-LSK114) run, basecalled with the model
dna_r10.4.1_e8.2_400bps_hac@v4.3.0 - One Assembly Polishing (SQK-APK114, using the 6b4 method) run, basecalled with a bespoke APK model that will be provided alongside the T2T bundle.
Data location
As with previous releases the new dataset is available for anonymous download from an Amazon Web Services S3 bucket. The bucket is part of the Open Data on AWS project enabling sharing and analysis of a wide range of data.
The data is located at:
s3://ont-open-data/londoncalling2024/assembly/
See the tutorials page for information on downloading datasets from S3.
The structure of the S3 prefix is shown below.
The primary assembly outputs are present under the assm prefix.
The basecalling outputs have been converted for to BAM for consistency using samtools import where there were not already present in this format.
The APK and and ULK reads aligned to the v1.0.1 HG002 T2T assembly (see below) are present in the qc-demo prefix.
Finally, a copy of this reference and a corresponding minimap2 index are under the ref prefix.
.├── assm│ ├── all_correct.fasta.gz│ ├── assembly.fasta│ ├── assembly.homopolymer-compressed.gfa│ └── medaka-6b4.fastq.gz├── basecalling│ ├── apk│ │ └── PAW41746.bam│ ├── pore-c│ │ └── PAW44788.bam│ └── ulk│ ├── PAW42495.bam│ └── PAW42666.bam├── qc-demo│ ├── apk-align│ │ ├── PAW41746.bam│ │ ├── PAW41746.bam.bai│ │ ├── PAW41746.histograms│ │ └── PAW41746.stats.gz│ └── ulk-align│ ├── PAW42495.bam│ ├── PAW42495.bam.bai│ ├── PAW42495.flagstats│ ├── PAW42495.histograms│ ├── PAW42495.stats.gz│ ├── PAW42666.bam│ ├── PAW42666.bam.bai│ ├── PAW42666.flagstats│ ├── PAW42666.histograms│ └── PAW42666.stats.gz├── ref│ ├── hg002v1.0.1.fasta.gz│ └── hg002v1.0.1.fasta.gz.mmi
Basecalling and QC analysis
Basecalling was performed using research-grade bonito, though results would be comparable with the newest version of dorado supporting transformer basecalling (v5.0.0).
For the purposes of exposition of data quality only, reads have been aligned to the v1.0.1 HG002 T2T assembly from the Telomere-to-Telomere Consortium. Alignment was performed for the ULK and APK datasets only, using the -x lr:hq preset of minimap2. Alignment statistics were calculated using the bamstats program available in the fastcat package.
Assembly workflow
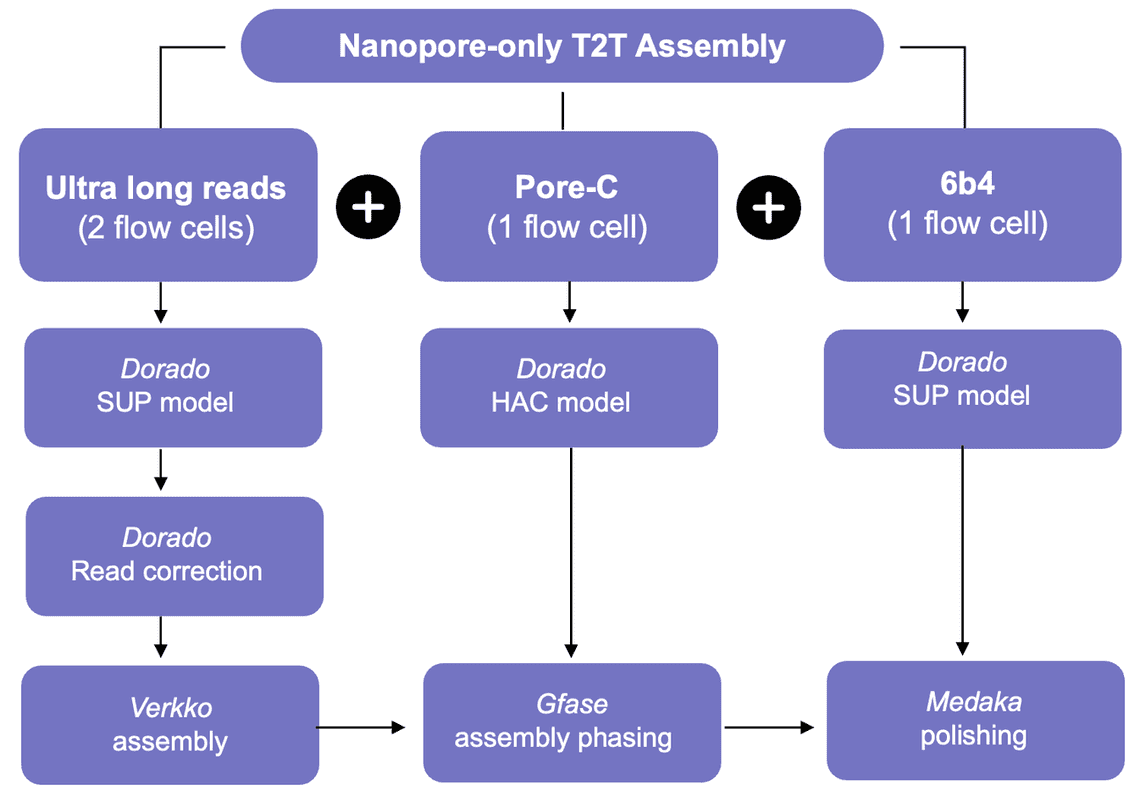
The Nanopore-only telomere-to-telomere assembly workflow is summarised in Figure 2.
Ultra Long (SQK-ULK114) reads pass through the self-correction algorithm now available in dorado.
This algorithm used is derived from that available in the herro project.
The corrected reads are then assembled using the verkko assembler. The assembler is run until the rukki step, at which data from Pore-C is injected into the verkko pipeline with GFAse being run to incorporate the Pore-C data into the assembly. The verkko pipeline is resumed by first re-running rukki and then continuing as normal. Finally medaka is run to correct the assembly further using the data from APK sequencing.
This workflow is embodied in the following prototype bash snippets.
It is assumed all the tools are available in the user’s environment. We begin by defining some files and paths and running verkko until the rukki step:
# set paths to relevant filesexport GFASE_DIR=<gfase install dir>export POREC_FASTA=<path to porec reads>export CORRECTED_READS=<path to corrected ulk reads>export RAW_READS=<path to uncorrected ulk reads># run verkko first time, up until rukki stepverrko --hifi "${CORRECTED_READS}" --nano "${RAW_READS}" \--hap-kmers /dev/null /dev/null trio -d verkko_results \--snakeopts "--until rukki"
We then incorporate the Pore-C sequencing data. This must be homopolymer compressed and then aligned to the unitigs created by verkko:
export MINIMAP_THREADS=256# homopolymer compress pore-c fastaseqtk hpc $POREC_FASTA > porec_simplex.hpc.fasta# align pore-c datapushd verkko_results/6-rukkiminimap2 -ax lr:hq -t ${MINIMAP_THREADS} -I 20G \unitig-unrolled-unitig-unrolled-popped-unitig-normal-connected-tip.fasta \../../porec_simplex.hpc.fasta \| samtools view -b@20 -q 1 \> porecVedges.bam
and phase the assembly graph using GFAse:
export GFASE_THREADS=48# run gfase$GFASE_DIR/build/phase_contacts_with_monte_carlo -i porecVedges.bam \-g ../5-untip/unitig-unrolled-unitig-unrolled-popped-unitig-normal-connected-tip.gfa \-o GFase --skip_unzip --use_homology -t ${GFASE_THREADS} -m 2
After this we need to do a little wrangling to obtain files from which we can continue the verkko workflow:
# convert gfase outputs to rukki inputsmv unitig-unrolled-unitig-unrolled-popped-unitig-normal-connected-tip.colors.csv \trio.colors.csvhead -n 1 trio.colors.csv > gfase.colors.csvcat GFase/phases.csv | awk 'BEGIN{FS=",";OFS="\t"} \$2=="1"&&$3>=10{print $1,$3*10,"0",$3*10":0","#FF8888"} \$2=="-1"&&$3>=10{print $1,"0",$3*10,"0:"$3*10,"#8888FF"} \$2=="1"&&$3<10{print $1,$3*10,"0",$3*10":0","#AAAAAA"} \$2=="-1"&&$3<10{print $1,"0",$3*10,"0:"$3*10,"#AAAAAA"}' \>> gfase.colors.csvawk '{print $1"\t"}' gfase.colors.csv > gfasedNodes.listfgrep -f gfasedNodes.list -v \unitig-unrolled-unitig-unrolled-popped-unitig-normal-connected-tip.noseq.gfa \| awk '/^S/{print $2"\t0\t0\t0:0\t#AAAAAA"}' \>> gfase.colors.csvcp gfase.colors.csv \unitig-unrolled-unitig-unrolled-popped-unitig-normal-connected-tip.colors.csv# jump back up to the top-level directorypopd
With that minor detour out of the way, we can continue our journey and finish off the verkko pipeline:
verrko --hifi "${CORRECTED_READS}" --nano "${RAW_READS}" \--hap-kmers /dev/null /dev/null trio -d verkko_results
Having run verkko in its entirety, the last step is to polish the assembly using data from the APK sequencing kit. We do this using medaka and a special consensus model that simultaneously uses both ULK and APK data for error correction:
APK_BAM=<path to apk reads as unaligned BAM>ULK_BAM=<path to ulk reads as unaligned BAM>VERKKO_ASSEMBLY=<path to verkko assembly>OUTPUT=assm-correctedmedaka_consensus_joint \-i "${APK_BAM}" -v apk -i "${ULK_BAM}" -v ulk \-t ${THREADS} -o "${OUTPUT}" \-m r1041_e82_260bps_joint_apk_ulk_v5.0.0 \-d "${VERKKO_ASSEMBLY}"
Further information
To register your interest in the T2T workflow, please click here. For additional information regarding these data please contact support@nanoporetech.com.
We hope that these data and analyses provide a useful resource to the community.
