Influenza Workflow
We are pleased to offer a new workflow for the analysis of targeted Oxford Nanopore Technologies sequencing of Influenza virus.
Influenza is a single stranded RNA virus and contains a 13.5-14.5kb genome which is split into 8 segments encoding 10-14 proteins (dependent on strain).
The virus is classified using two proteins found on the outer surface of the viral capsid. You’ve probably heard of H1N1 Influenza for example. The H represents hemagglutinin and the N is neuraminidase.
The Oxford Nanopore Technologies protocol listed here amplifies segments of the Influenza Type A and Type B genomes. Using the analysis workflow described here users can determine the most likely strain of Influenza to which the sample being sequenced belongs.
As with all our workflows we welcome your comments and suggestions for improvements or any reports of isssues on the nanopore community or on GitHub.
How To Run
You have two options for local analysis of your Influenza data.
1. Command Line
nextflow nextflow run epi2me-labs/wf-flu --fastq <PATH_TO_DEMULTIPLEXED_FASTQS>
Optional parameters include
--sample_sheet <PATH_TO_SAMPLESHEET>(Example below)--downsample <NUMBER_OF_READS>(default: None, suggested: 500)--min_qscore <MINIMUM_READ_QSCORE>(default: 9)
Sample Sheet
A sample sheet allows you to name your samples, it must be a comma or tab separated file like below (note: this is equivalent to that which can be provided to MinKNOW, the workflow makes use of only a subset of the columns as shown below):
barcode,sample_id,typebarcode02,H1N1_strain_A-PR-8-34,test_samplebarcode03,H1N1_strain_A-Virginia-ATCC1-2009,test_samplebarcode04,H3N2_strain_A-Virginia-ATCC6-2012,test_samplebarcode08,fluB-BY-massachusetts-2-2012,test_samplebarcode09,fluB_B-taiwan-2-62,test_samplebarcode10,fluB-lee-40,test_samplebarcode31,fluB-yamagata-florida-4-2006_1,test_samplebarcode91,fluB-yamagata-florida-4-2006_2,test_samplebarcode55,fluA_H3N2_A-wisconsin-15-2009,test_sample
2. EPI2ME Labs
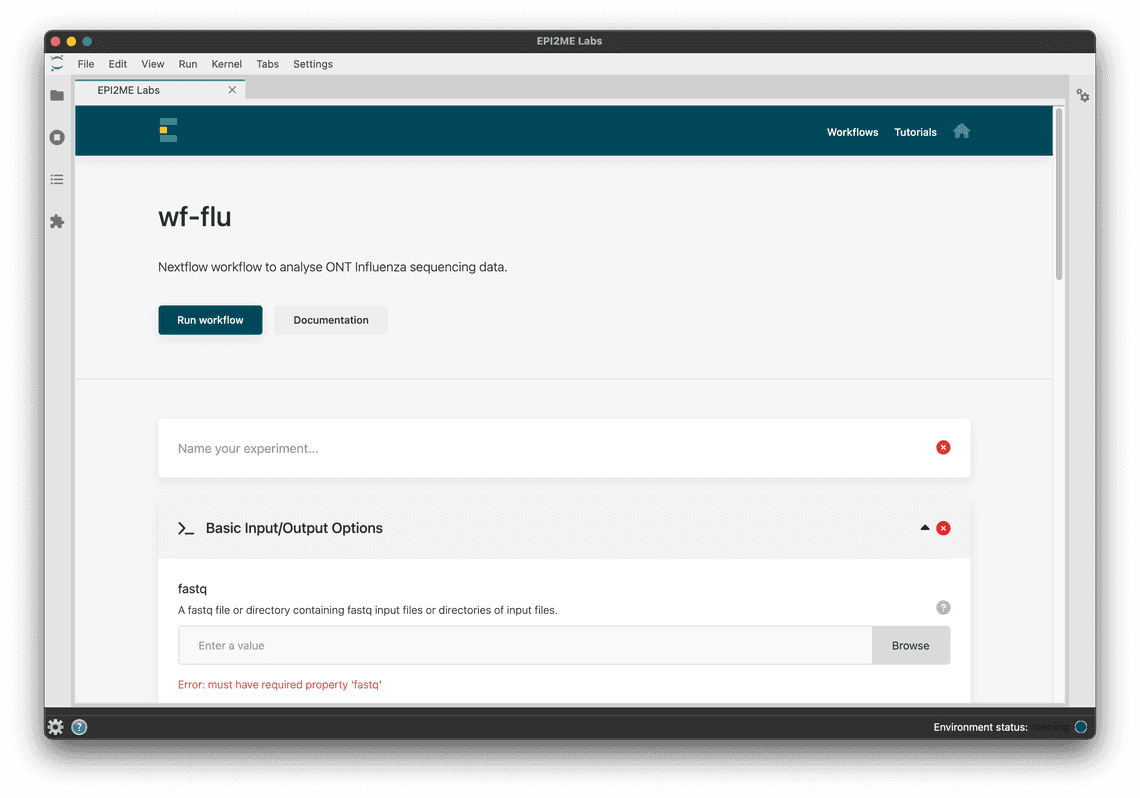
You can also use EPI2ME Labs, our straightforward application for the point and click execution of Nextflow workflows. This is available for Windows, MacOS and ubuntu on our downloads page.
Data Analysis Details
The workflow analyses all samples on a multiplexed Influenza sequencing run and provide an easy to interpret report.
The workflow carries out the following steps:
- Concatenate reads from the same sample & filter out short reads < 200 bases long
- Filter reads below median qscore of 9 (can be adjusted by changing
--min_qscore parameter) - Align reads to reference (minimap2)
- Optional downsampleing to speed up execution (activate by using
--downsample 500) - Coverage calculations (samtools)
- Call variants with medaka
- Make a (coverage masked) consensus with bcftools
- Perform Influenza typing with abricate
Reference
We align to the CDC’s multi-fasta Influenza reference which contains FluA + FluB segments, as well as including alternative segment sequences from disparate strains:
A_MP, A_NP, A_NS, A_PA, A_PB1, A_PB2, A_HA_H1, A_HA_H10, A_HA_H11, A_HA_H12, A_HA_H13, A_HA_H14, A_HA_H15, A_HA_H16, A_HA_H2, A_HA_H3, A_HA_H4, A_HA_H5, A_HA_H6, A_HA_H7, A_HA_H8, A_HA_H9, A_NA_N1, A_NA_N2, A_NA_N3, A_NA_N4, A_NA_N5, A_NA_N6, A_NA_N7, A_NA_N8, A_NA_N9, B_HA, B_MP, B_NA, B_NP, B_NS, B_PA, B_PB1, B_PB2
Downsampling
Downsampling is optional and will speed up workflow execution.
For every segment in the reference genome, the workflow:
- Finds the length of the segment
- Finds reads within ±10% of the segment length
- Collates a balanced set of reads from forward and reverse strands to achieve the desired read count
Typing
Typing is carried out using abricate with the INSaFLU database containing the sequences in the table below.
| Database | Gene | Accession | Details |
|---|---|---|---|
| insaflu | M1 | MK576795 | Type_A MK576795 A/England/7821/2019 2019/01/03 7 (MP) |
| insaflu | M1 | AF100378 | Type_B AF100378.1 Influenza B virus B/Yamagata/16/88 segment 7 M1 matrix protein (M) and BM2 protein (BM2) genes, complete cds |
| insaflu | HA | FJ966974 | H1 FJ966974.1 Influenza A virus (A/California/07/2009(H1N1)) segment 4 hemagglutinin (HA) gene, complete cds |
| insaflu | HA | L11142 | H2 L11142.1 Influenza A virus (A/Singapore/1/57 (H2N2)) hemagglutinin (HA) gene, complete cds |
| insaflu | HA | MK576794 | H3 MK576794 A/England/7821/2019 2019/01/03 4 (HA) |
| insaflu | HA | AF285883 | H4 AF285883.2 Influenza A virus (A/Swine/Ontario/01911-2/99 (H4N6)) segment 4 hemagglutinin (HA) gene, complete cds |
| insaflu | HA | EF541403 | H5 EF541403.1 Influenza A virus (A/Viet Nam/1203/2004(H5N1)) segment 4 hemagglutinin (HA) gene, complete cds |
| insaflu | HA | AB295613 | H15 AB295613.1 Influenza A virus (A/duck/Australia/341/83(H15N8)) HA gene for haemagglutinin, complete cds |
| insaflu | NA | GQ377078 | N1 GQ377078.1 Influenza A virus (A/California/07/2009(H1N1)) segment 6 neuraminidase (NA) gene, complete cds |
| insaflu | NA | MK576796 | N2 MK576796 A/England/7821/2019 2019/01/03 6 (NA) |
| insaflu | NA | AB295614 | N8 AB295614.1 Influenza A virus (A/duck/Australia/341/83(H15N8)) NA gene for neuraminidase, complete cds |
| insaflu | HA | AY338459 | H7 AY338459.1 Influenza A virus (A/Netherlands/219/2003(H7N7)) segment 4 hemagglutinin (HA) gene, complete cds |
| insaflu | HA | CY014659 | H8 CY014659.1 Influenza A virus (A/turkey/Ontario/6118/1968(H8N4)) segment 4, complete sequence |
| insaflu | HA | CY014694 | H13 CY014694.1 Influenza A virus (A/gull/Maryland/704/1977(H13N6)) segment 4, complete sequence |
| insaflu | HA | CY018765 | Yamagata CY018765.1 Influenza B virus (B/Yamagata/16/1988) segment 4, complete sequence |
| insaflu | HA | CY103892 | H17 CY103892.1 Influenza A virus (A/little yellow-shouldered bat/Guatemala/060/2010(H17N10)) hemagglutinin (HA) gene, complete cds |
| insaflu | NA | CY103894 | N10 CY103894.1 Influenza A virus (A/little yellow-shouldered bat/Guatemala/060/2010(H17N10)) neuraminidase (NA) gene, complete cds |
| insaflu | NA | CY125730 | N3v2 CY125730.1 Influenza A virus (A/Mexico/InDRE7218/2012(H7N3)) neuraminidase (NA) gene, complete cds |
| insaflu | HA | CY125945 | H18 CY125945.1 Influenza A virus (A/flat-faced bat/Peru/033/2010(H18N11)) hemagglutinin (HA) gene, complete cds |
| insaflu | NA | CY125947 | N11 CY125947.1 Influenza A virus (A/flat-faced bat/Peru/033/2010(H18N11)) neuraminidase-like protein (NA) gene, complete cds |
| insaflu | HA | CY130078 | H12 CY130078.1 Influenza A virus (A/duck/Alberta/60/1976(H12N5)) hemagglutinin (HA) gene, complete cds |
| insaflu | HA | CY130094 | H14 CY130094.1 Influenza A virus (A/mallard/Astrakhan/263/1982(H14N5)) hemagglutinin (HA) gene, complete cds |
| insaflu | NA | CY130096 | N5 CY130096.1 Influenza A virus (A/mallard/Astrakhan/263/1982(H14N5)) neuraminidase (NA) gene, complete cds |
| insaflu | HA | DQ376624 | H6 DQ376624.1 Influenza A virus (A/chicken/Taiwan/0705/99(H6N1)) hemagglutinin (HA) gene, complete cds |
| insaflu | HA | EU293864 | H16 EU293864.1 Influenza A virus (A/black-headed gull/Turkmenistan/13/76(H16N3)) hemagglutinin (HA) gene, complete cds |
| insaflu | HA | FJ183474 | H10 FJ183474.1 Influenza A virus (A/mallard/Bavaria/3/2006(H10N7)) segment 4 hemagglutinin (HA) gene, complete cds |
| insaflu | NA | FJ183475 | N7 FJ183475.1 Influenza A virus (A/mallard/Bavaria/3/2006(H10N7)) segment 6 neuraminidase (NA) gene, complete cds |
| insaflu | NA | GQ907296 | N3v1 GQ907296.1 Influenza A virus (A/black headed gull/Mongolia/1756/2006(H16N3)) segment 6 neuraminidase (NA) gene, complete cds |
| insaflu | HA | GU052203 | H11 GU052203.1 Influenza A virus (A/duck/England/1/1956(H11N6)) segment 4 hemagglutinin (HA) gene, complete cds |
| insaflu | NA | KC853765 | N9 KC853765.1 Influenza A virus (A/Hangzhou/1/2013(H7N9)) segment 6 neuraminidase (NA) gene, complete cds |
| insaflu | HA | KX879589 | H9 KX879589.1 Influenza A virus (A/swine/Hong Kong/9/98(H9N2)) segment 4 hemagglutinin (HA) gene, partial cds |
| insaflu | HA | M58428 | Victoria M58428.1 Influenza B/Victoria/2/87, hemagglutinin (seg 4), RNA |
| insaflu | NA | EU429793 | N4 EU429793.1 Influenza A virus (A/turkey/Ontario/6118/1968(H8N4)) segment 6 neuraminidase (NA) mRNA, complete cds |
| insaflu | NA | EU429795 | N6 EU429795.1 Influenza A virus (A/duck/England/1/1956(H11N6)) segment 6 neuraminidase (NA) mRNA, complete cds |
Output Files
The workflow outputs several files that are useful for interpretation and analysis:
- Per run:
wf-flu-report.html: Easy to use HTML report for all samples on the runwf-flu-results.csv: Typing results in CSV format for onward processing
- Per sample:
<SAMPLE_NAME>.stats: Read stats<SAMPLE_NAME>.bam: Alignment of reads to reference<SAMPLE_NAME>.bam.bai: BAM index<SAMPLE_NAME>.annotate.filtered.vcf: medaka called variants<SAMPLE_NAME>.draft.consensus.fasta: Consensus FASTA<SAMPLE_NAME>.insaflu.typing.txt: abricate typing results<SAMPLE_NAME>.depth.txt: samtools depth, columns are contig, postion, and coverage
Useful Links
- INSaFLU: https://insaflu.insa.pt/
Share
Table Of Contents
Related Posts
Information

