Importing third-party workflows into EPI2ME Labs

With the release of EPI2ME Labs version 4.0.0 we are now able to run community developed Nextflow workflows from the application’s interface. Users unfamiliar with a command line environment can benefit from cutting-edge open source bioinformatic analysis developed by the community.
If you are a developer of Nextflow workflows for the analysis of Oxford Nanopore sequencing data, then following a few simple recommendations will provide the best possible user experience.
EPI2ME Labs style guide
We believe that a workflow should take users from standard sequencing instrument outputs to rich outputs that answer a scientific question or allow users to formulate new ones.
Throwing the kitchen sink at a workflow leads to a complex set of parameters for a user to understand and a steep learning curve which might put off those prospective users. Our suggestion is that developers do not overload their workflows with a scatter gun approach to tool inclusion, and try to keep to a single theme.
We’d like to suggest three simple rules to guide developers:
- Don’t try to be everything to everyone,
- Provide useful outputs,
- Notwithstanding rule 1, modularise functionality and make any discrete components optional,
Don’t try to be everything to everyone
We recommend creating workflows based on a single end-user analysis theme or application. For example we would advise against making a workflow to “analyse Nanopore data”. It is too easy for workflow developers to become obsessed with including more and more tools and features.
Your scientists were so preoccupied with whether they could, they didn’t stop to think if they should
— Dr. Ian Malcolm, Jurassic Park
Rather developers should create workflows that solve a data analysis problem. Several EPI2ME Labs workflows provide a very narrow range of functionality or service a particular application; our wf-clone-validation services the needs of users who wish to verify the sequence of their artificially contructed plasmid sequences. It bears no shame in doing one thing well.
Provide meaningful outputs
Workflows typically run a variety of bioinformatics tools. Each tool may output a multitude of files. Not all of these will be useful to end-users and publishing all conceivable outputs can only lead to confusion. We recommend publishing only those outputs in line with the ultimate analysis aims of the workflow.
In addition to providing the primary analysis results, we recommend summarising results in a consolidated fashion. All EPI2ME Labs workflows create a HTML document including rich plots.
Modularise functionality
Having decided on an analysis theme, we would still recommend making discrete functionality optional. Our own wf-human-variation workflow can provide a variety of analyses all on the theme of analysing genetic variation in human samples. It provides small variant calling, structural variants, modified base analysis and forthcoming releases will provide information on copy number variation and tandem repeats. Each of these operates independently such that users can mix and match which components are run, appropriate to their scientific question.
The nextflow.config
The EPI2ME Labs user interface generates much of its content from both the nextflow.config
file and the nextflow_schema.json file, which should therefore be present in all
workflow projects.
In our own workflows we have recently dropped support for conda as a means of defining and distributing software dependencies. Users reported many issues with use of conda in their environments; it can be described as temperamental at best.
Our workflows by default use Docker to provide their software components. We also define a singularity profile for users who are unable to use Docker. You will see that our workflows also contain a parameterised stub for use of AWS batch — this is really only for our own selfish convenience, but can also be used be users to intergrate with AWS services.
On the topic of Docker, and related to why its not an option for many, we highly recommend creating Docker images that do not run as the root user by default. Instead workflow authors should prepare images that run as a standard user in a group. Having done this Nextflow can be instructed to use this user in any containers that it creates. We set also some basic options for singularity users:
standard {docker {enabled = true// this ensures container is run as host user and group, but// also adds host user to the within-container grouprunOptions = "--user \$(id -u):\$(id -g) --group-add 100"}}singularity {singularity {enabled = trueautoMounts = true}}
The configuration files should contain also some basic information in its manifest. Items here are used by EPI2ME Labs to help the user identify workflows and their versions:
manifest {name = 'epi2me-labs/wf-tb-amr'author = 'Oxford Nanopore Technologies'homePage = 'https://github.com/epi2me-labs/wf-tb-amr'description = 'Anti-microbial agent resistance calling for Mycobacterium tuberculosis'mainScript = 'main.nf'nextflowVersion = '>=20.10.0'version = 'v1.0.11'}
EPI2ME Labs makes heavy use of the JSON schema parameter specification developed by nf-core. The schema drives the creation of the workflow execution screen of EPI2ME Labs 4.0.0. It is therefore important that this file follows valid JSON schema syntax and conventions of the nf-core project.
For the user interface of EPI2ME Labs to display useful input form components
the type of each paramter should be correct. For example specifying path
will render a file or directory selection input box, while file-path will
render a dialog where only files can be selected. Similarly use of enum
will lead to the UI rendering a drop-down value selector. This is preferable
to a free text input for parameters that accept only a discrete range of
values.
To give useful prompts to the user we encourage the use of a short but
informative description, and more verbose help_text, for every parameter
in the schema. In our own development we have found it useful to have developers
not familiar with the project to provide input and feedback on the parameter
descriptions. What can be obvious to the primary developer of a workflow can
be utterly opaque to even knowledgeable co-workers.
These descriptions and help texts should therefore be written assuming the the user is unfamilar with both your workflow and the parameter in question. Guidance should be provided on when and why a parameter might want to be changed.
As an example parameter entry, the following is the FASTQ file specification for almost all EPI2ME Labs workflows:
"fastq": {"type": "string","format": "path","description": "FASTQ files to use in the analysis.","help_text": "This accepts one of three cases: (i) the path to a single FASTQ file; (ii) the path to a top-level directory containing FASTQ files; (iii) the path to a directory containing one level of sub-directories which in turn contain FASTQ files. In the first and second case, a sample name can be supplied with `--sample`. In the last case, the data is assumed to be multiplexed with the names of the sub-directories as barcodes. In this case, a sample sheet can be provided with `--sample_sheet`."}
Finally we highly recommend the use of meaningful parameter groups to help orient users. All EPI2ME Labs workflows have minimally:
Input Options- Parameters related to input data, we use this for the location of FASTQ files, for example.Sample Options- Parameters related to the samples being analysed, we use this for the location of a sample sheet, for example.Output Options- Parameters related to the output of the results of the workflow.Advanced Options- Options that an everyday user perhaps wouldn’t need to interact with.Miscellaneous Options- All the other stuff.
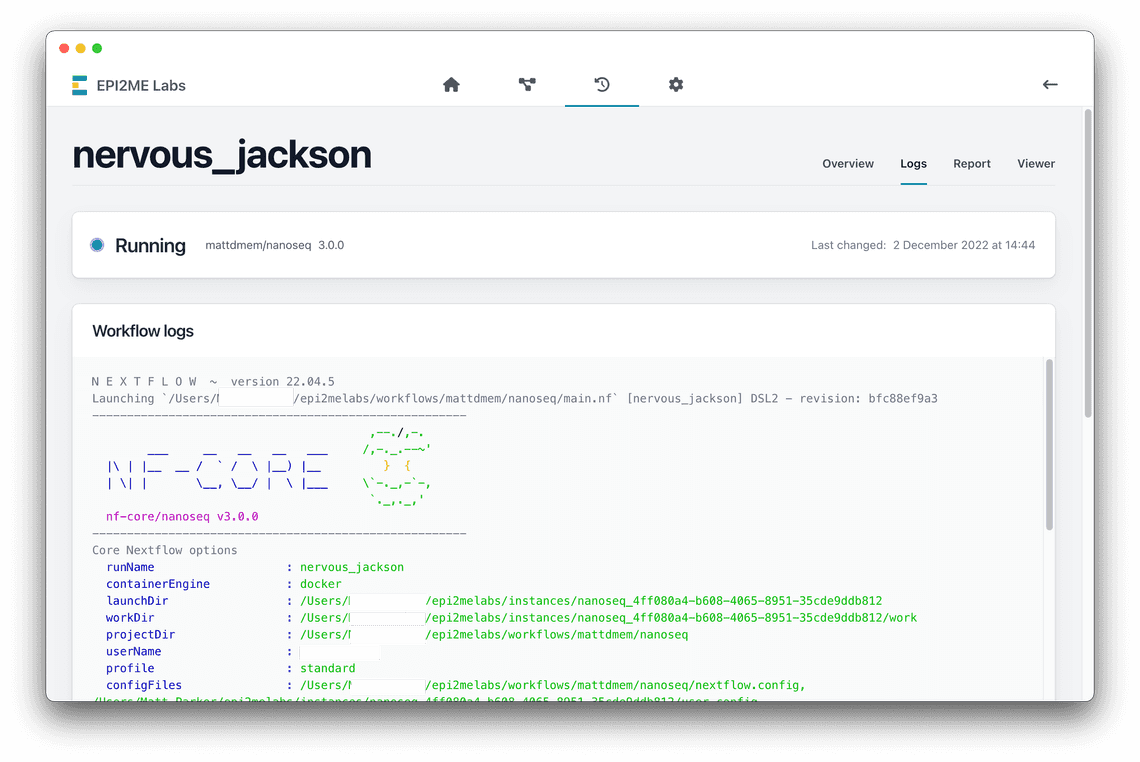
Importing nanoseq
As an example of importing a third party workflow into EPI2ME Labs, the
nanoseq workflow can be used by making the following adjustments, some of
which fix errors in the original:
- The parameter for the selection of DNA/RNA/cDNA was wrong in the original schema issues/203
- We prefer to skip most of the complex options in the workflow by default. The workflow can perform basecalling, demultiplexing, transcriptomic analysis and a host of other unrelated analyses.
- We’ve included a
demo_urlwhich links to a demonstration dataset for users to test the workflow. - The default compute requirements were prohibitive for many compute environments; so we’ve reduced the default memory specification.
To use this workflow, click “Import workflow” on the workflows section of the app and enter the following URL
https://github.com/epi2me-labs/nanoseq
Click import and the workflow should download and appear as one of the workflows to select; the workflow can be run with the demonstration dataset by clicking “Use demo data”.
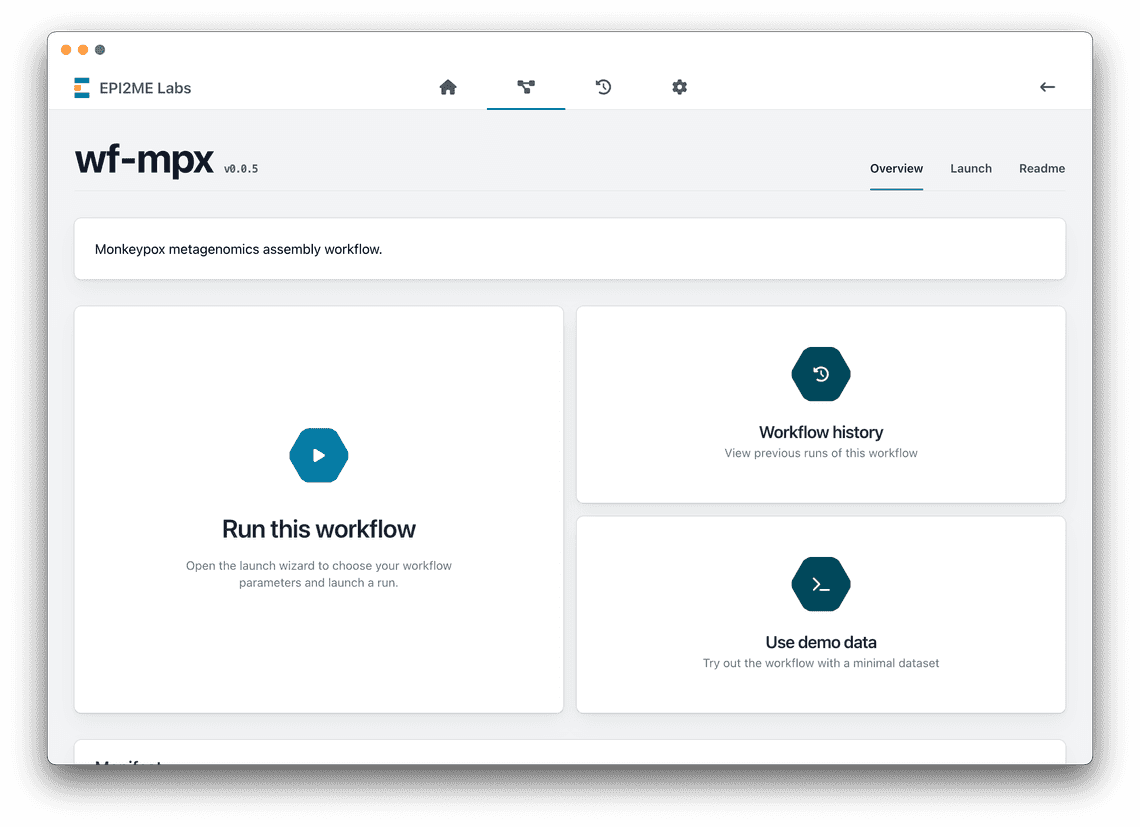
Adding demonstration data to your workflow
EPI2ME Labs includes a button that allows the user to run the workflow without having to first get to grips with the often complex parameters of a workflow. This is a great opportunity to demonstrate to a user what is achievable with your workflow, and the type of outputs they can expect. Workflows which would otherwise be valuable to the community go unused when they suffer from poor user interface and experience.
We suggest that demo data should include a dataset that show cases all the features of your workflow, whilst being quick to run.
Our own demo datasets are structured along the following lines:
my-workflow-demo├── fastq│ └── barcode01│ └── reads.fastq.gz└── nextflow.config
The directory structure is self-contained, with a nextflow.config with all necessary parameters for the workflow to run successfully without the user having to provide additional parameters to nextflow. (If you find yourself specifying many parameters in your demo nextflow configuration file, that may suggest that you need to amend the underlying workflow).
With the structure above we suggest placing an archive of the entire contents on a publicly accessible location. Our own datasets are stored in Amazon Web Services S3 buckets, accessible through the AWS CLI or more plainly with any HTTP client. As an extension to the standard nf-core parameter schemas, EPI2ME Labs can parse a special top-level key demo_url containing a URL to fetch a compressed archive:
"demo_url": "https://public-url-to-my-bucket/my-workflow/my-workflow-demo.tar.gz"
wf-template
For those wanting a particularly easy way to get started, our template workflow wf-template encodes all of the above practices into a simple, yet functional, basic workflow. The workflow doesn’t really do much in terms of analysis but it contains all the library code that we use for data ingress and implements all of the patterns noted above. The project can be forked and immediately used and extended.
Summary
Nexflow workflows developed by the EPI2ME Labs team at Oxford Nanopore follow a minimal template driven by and focussed on creating the best possible experience for end users. We focus on ease of use and answering the scientific questions of researchers.
The EPI2ME Labs interface has been designed to complement this ethos of usability and productivity. EPI2ME Labs builds on existing community standards: users are able to interact with Nextflow workflows developed by third parties not just those from Oxford Nanopore
We invite workflow developers to download EPI2ME Labs and test the integration of their workflows and reach a wider audience.
Share
Table Of Contents
Related Posts
Information

